| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Detected and Quantified |
|---|
| Creation Date | 2005-11-16 15:48:42 UTC |
|---|
| Update Date | 2021-09-14 14:58:04 UTC |
|---|
| HMDB ID | HMDB0001248 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
| Common Name | FAD |
|---|
| Description | Flavin adenine dinucleotide (FAD) is a redox-active coenzyme associated with various proteins, which is involved with several enzymatic reactions in metabolism. FAD, also known as adeflavin or flamitajin b, belongs to the class of organic compounds known as flavin nucleotides. These are nucleotides containing a flavin moiety. Flavin is a compound that contains the tricyclic isoalloxazine ring system, which bears 2 oxo groups at the 2- and 4-positions. FAD is a drug which is used to treat eye diseases caused by vitamin b2 deficiency, such as keratitis and blepharitis. FAD exists in all living species, ranging from bacteria to humans. In humans, FAD is involved in the metabolic disorder called the medium chain acyl-coa dehydrogenase deficiency (mcad) pathway. Outside of the human body, FAD has been detected, but not quantified in several different foods, such as other bread, passion fruits, asparagus, kelps, and green bell peppers. It is a flavoprotein in which the substituent at position 10 of the flavin nucleus is a 5'-adenosyldiphosphoribityl group. |
|---|
| Structure | CC1=CC2=C(C=C1C)N(C[C@H](O)[C@H](O)[C@H](O)COP(O)(=O)OP(O)(=O)OC[C@H]1O[C@H]([C@H](O)[C@@H]1O)N1C=NC3=C1N=CN=C3N)C1=NC(=O)NC(=O)C1=N2 InChI=1S/C27H33N9O15P2/c1-10-3-12-13(4-11(10)2)35(24-18(32-12)25(42)34-27(43)33-24)5-14(37)19(39)15(38)6-48-52(44,45)51-53(46,47)49-7-16-20(40)21(41)26(50-16)36-9-31-17-22(28)29-8-30-23(17)36/h3-4,8-9,14-16,19-21,26,37-41H,5-7H2,1-2H3,(H,44,45)(H,46,47)(H2,28,29,30)(H,34,42,43)/t14-,15+,16+,19-,20+,21+,26+/m0/s1 |
|---|
| Synonyms | | Value | Source |
|---|
| Adenosine 5'-(trihydrogen pyrophosphate), 5'-5'-ester with riboflavine | ChEBI | | Adenosine 5'-[3-(riboflavin-5'-yl) dihydrogen diphosphate] | ChEBI | | Flavin adenine dinucleotide | ChEBI | | FLAVIN-adenine dinucleotide | ChEBI | | Riboflavin 5'-(trihydrogen diphosphate), 5'-5'-ester with adenosine | ChEBI | | Riboflavin 5'-adenosine diphosphate | ChEBI | | Adeflavin | Kegg | | Adenosine 5'-(trihydrogen pyrophosphoric acid), 5'-5'-ester with riboflavine | Generator | | Adenosine 5'-[3-(riboflavin-5'-yl) dihydrogen diphosphoric acid] | Generator | | Riboflavin 5'-(trihydrogen diphosphoric acid), 5'-5'-ester with adenosine | Generator | | Riboflavin 5'-adenosine diphosphoric acid | Generator | | 1H-Purin-6-amine flavin dinucleotide | HMDB | | 1H-Purin-6-amine flavine dinucleotide | HMDB | | Adenine-flavin dinucleotide | HMDB | | Adenine-flavine dinucleotide | HMDB | | Adenine-riboflavin dinuceotide | HMDB | | Adenine-riboflavin dinucleotide | HMDB | | Adenine-riboflavine dinucleotide | HMDB | | Flamitajin b | HMDB | | Flanin F | HMDB | | Flavin adenine dinucleotide oxidized | HMDB | | Flavine adenosine diphosphate | HMDB | | Flavine-adenine dinucleotide | HMDB | | Flavitan | HMDB | | Flaziren | HMDB | | Isoalloxazine-adenine dinucleotide | HMDB | | Riboflavin-adenine dinucleotide | HMDB | | Riboflavine-adenine dinucleotide | HMDB | | Dinucleotide, flavin-adenine | HMDB |
|
|---|
| Chemical Formula | C27H33N9O15P2 |
|---|
| Average Molecular Weight | 785.5497 |
|---|
| Monoisotopic Molecular Weight | 785.157134455 |
|---|
| IUPAC Name | {[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-3,4-dihydroxyoxolan-2-yl]methoxy}[({[(2R,3S,4S)-5-{7,8-dimethyl-2,4-dioxo-2H,3H,4H,10H-benzo[g]pteridin-10-yl}-2,3,4-trihydroxypentyl]oxy}(hydroxy)phosphoryl)oxy]phosphinic acid |
|---|
| Traditional Name | flavine-adenine dinucleotide |
|---|
| CAS Registry Number | 146-14-5 |
|---|
| SMILES | CC1=CC2=C(C=C1C)N(C[C@H](O)[C@H](O)[C@H](O)COP(O)(=O)OP(O)(=O)OC[C@H]1O[C@H]([C@H](O)[C@@H]1O)N1C=NC3=C1N=CN=C3N)C1=NC(=O)NC(=O)C1=N2 |
|---|
| InChI Identifier | InChI=1S/C27H33N9O15P2/c1-10-3-12-13(4-11(10)2)35(24-18(32-12)25(42)34-27(43)33-24)5-14(37)19(39)15(38)6-48-52(44,45)51-53(46,47)49-7-16-20(40)21(41)26(50-16)36-9-31-17-22(28)29-8-30-23(17)36/h3-4,8-9,14-16,19-21,26,37-41H,5-7H2,1-2H3,(H,44,45)(H,46,47)(H2,28,29,30)(H,34,42,43)/t14-,15+,16+,19-,20+,21+,26+/m0/s1 |
|---|
| InChI Key | VWWQXMAJTJZDQX-UYBVJOGSSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as flavin nucleotides. These are nucleotides containing a flavin moiety. Flavin is a compound that contains the tricyclic isoalloxazine ring system, which bears 2 oxo groups at the 2- and 4-positions. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Nucleosides, nucleotides, and analogues |
|---|
| Class | Flavin nucleotides |
|---|
| Sub Class | Not Available |
|---|
| Direct Parent | Flavin nucleotides |
|---|
| Alternative Parents | |
|---|
| Substituents | - Flavin nucleotide
- (3'->5')-dinucleotide
- (3'->5')-dinucleotide or analogue
- Purine ribonucleoside diphosphate
- Purine ribonucleoside monophosphate
- Flavin
- Isoalloxazine
- Pentose-5-phosphate
- Pentose phosphate
- Glycosyl compound
- N-glycosyl compound
- Diazanaphthalene
- Pentose monosaccharide
- Pteridine
- 6-aminopurine
- Quinoxaline
- Organic pyrophosphate
- Monosaccharide phosphate
- Imidazopyrimidine
- Purine
- Monoalkyl phosphate
- Aminopyrimidine
- Pyrimidone
- Monosaccharide
- N-substituted imidazole
- Organic phosphoric acid derivative
- Imidolactam
- Benzenoid
- Alkyl phosphate
- Phosphoric acid ester
- Pyrimidine
- Pyrazine
- Tetrahydrofuran
- Azole
- Vinylogous amide
- Heteroaromatic compound
- Imidazole
- Secondary alcohol
- Lactam
- Polyol
- Oxacycle
- Azacycle
- Organoheterocyclic compound
- Organic oxide
- Organic oxygen compound
- Organopnictogen compound
- Organonitrogen compound
- Hydrocarbon derivative
- Primary amine
- Alcohol
- Amine
- Organic nitrogen compound
- Organooxygen compound
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Not Available | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | 5 mg/mL | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Experimental Chromatographic Properties | Experimental Collision Cross Sections |
|---|
| Predicted Molecular Properties | |
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Retention Times Underivatized| Chromatographic Method | Retention Time | Reference |
|---|
| Measured using a Waters Acquity ultraperformance liquid chromatography (UPLC) ethylene-bridged hybrid (BEH) C18 column (100 mm × 2.1 mm; 1.7 μmparticle diameter). Predicted by Afia on May 17, 2022. Predicted by Afia on May 17, 2022. | 3.18 minutes | 32390414 | | Predicted by Siyang on May 30, 2022 | 11.8294 minutes | 33406817 | | Predicted by Siyang using ReTip algorithm on June 8, 2022 | 9.26 minutes | 32390414 | | AjsUoB = Accucore 150 Amide HILIC with 10mM Ammonium Formate, 0.1% Formic Acid | 508.7 seconds | 40023050 | | Fem_Long = Waters ACQUITY UPLC HSS T3 C18 with Water:MeOH and 0.1% Formic Acid | 1423.7 seconds | 40023050 | | Fem_Lipids = Ascentis Express C18 with (60:40 water:ACN):(90:10 IPA:ACN) and 10mM NH4COOH + 0.1% Formic Acid | 215.2 seconds | 40023050 | | Life_Old = Waters ACQUITY UPLC BEH C18 with Water:(20:80 acetone:ACN) and 0.1% Formic Acid | 54.2 seconds | 40023050 | | Life_New = RP Waters ACQUITY UPLC HSS T3 C18 with Water:(30:70 MeOH:ACN) and 0.1% Formic Acid | 175.7 seconds | 40023050 | | RIKEN = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 78.0 seconds | 40023050 | | Eawag_XBridgeC18 = XBridge C18 3.5u 2.1x50 mm with Water:MeOH and 0.1% Formic Acid | 422.5 seconds | 40023050 | | BfG_NTS_RP1 =Agilent Zorbax Eclipse Plus C18 (2.1 mm x 150 mm, 3.5 um) with Water:ACN and 0.1% Formic Acid | 393.6 seconds | 40023050 | | HILIC_BDD_2 = Merck SeQuant ZIC-HILIC with ACN(0.1% formic acid):water(16 mM ammonium formate) | 707.1 seconds | 40023050 | | UniToyama_Atlantis = RP Waters Atlantis T3 (2.1 x 150 mm, 5 um) with ACN:Water and 0.1% Formic Acid | 664.9 seconds | 40023050 | | BDD_C18 = Hypersil Gold 1.9µm C18 with Water:ACN and 0.1% Formic Acid | 366.5 seconds | 40023050 | | UFZ_Phenomenex = Kinetex Core-Shell C18 2.6 um, 3.0 x 100 mm, Phenomenex with Water:MeOH and 0.1% Formic Acid | 827.0 seconds | 40023050 | | SNU_RIKEN_POS = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 202.8 seconds | 40023050 | | RPMMFDA = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 314.4 seconds | 40023050 | | MTBLS87 = Merck SeQuant ZIC-pHILIC column with ACN:Water and :ammonium carbonate | 696.6 seconds | 40023050 | | KI_GIAR_zic_HILIC_pH2_7 = Merck SeQuant ZIC-HILIC with ACN:Water and 0.1% FA | 344.3 seconds | 40023050 | | Meister zic-pHILIC pH9.3 = Merck SeQuant ZIC-pHILIC column with ACN:Water 5mM NH4Ac pH9.3 and 5mM ammonium acetate in water | 578.7 seconds | 40023050 |
Predicted Kovats Retention IndicesUnderivatized |
|---|
| Spectra |
|---|
| GC-MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (Non-derivatized) - 70eV, Positive | splash10-0170-1032960400-cdd3286ebb1dea7c4f54 | 2017-11-06 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (TMS_1_1) - 70eV, Positive | Not Available | 2021-10-18 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (TMS_1_2) - 70eV, Positive | Not Available | 2021-10-18 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (TMS_1_3) - 70eV, Positive | Not Available | 2021-10-18 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (TMS_1_4) - 70eV, Positive | Not Available | 2021-10-18 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (TMS_1_5) - 70eV, Positive | Not Available | 2021-10-18 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (TMS_1_6) - 70eV, Positive | Not Available | 2021-10-18 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (TMS_1_7) - 70eV, Positive | Not Available | 2021-10-18 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (TMS_1_8) - 70eV, Positive | Not Available | 2021-10-18 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (TMS_1_9) - 70eV, Positive | Not Available | 2021-10-18 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | 2021-10-18 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (TBDMS_1_2) - 70eV, Positive | Not Available | 2021-10-18 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (TBDMS_1_3) - 70eV, Positive | Not Available | 2021-10-18 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (TBDMS_1_4) - 70eV, Positive | Not Available | 2021-10-18 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (TBDMS_1_5) - 70eV, Positive | Not Available | 2021-10-18 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (TBDMS_1_6) - 70eV, Positive | Not Available | 2021-10-18 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (TBDMS_1_7) - 70eV, Positive | Not Available | 2021-10-18 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (TBDMS_1_8) - 70eV, Positive | Not Available | 2021-10-18 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - FAD GC-MS (TBDMS_1_9) - 70eV, Positive | Not Available | 2021-10-18 | Wishart Lab | View Spectrum |
MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD Quattro_QQQ 10V, Positive-QTOF (Annotated) | splash10-000i-0001200900-b0740b3d33d50996ccde | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD Quattro_QQQ 25V, Positive-QTOF (Annotated) | splash10-000m-0105900000-3abff26d5bb35f8527b8 | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD Quattro_QQQ 40V, Positive-QTOF (Annotated) | splash10-000i-0931700000-73f360589a13230eea7a | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-000j-0908600300-b1eccda8ebae8b9e4cc1 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-000i-0900000000-bdb826f9c3cbc09eff9b | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-00di-0019800000-00ad56b6b6a3bb4516e6 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-000b-0009400000-667064ca470a5c341974 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-000j-0509700500-8766cb874f927ed5a795 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-000i-0900000000-3125e04c09a14c62f22a | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-00di-0019700000-78c46ea4b4f562757e56 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-000b-0009400000-94b845d2c8d082b9484c | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-001i-0000100900-6bac7b7f631dfc074a0c | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-001i-0920000000-49c9d4fb9b57a59f45a5 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-000i-0003900000-572f0b1bd59f71ab3c5f | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-000i-0003900000-d2ede6a2e7183f1c32a3 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-001i-0000100900-78554afbc26abe26af35 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-001i-0930000000-274da0a01c651b791733 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-000i-0003900000-55939cda9e14fc582757 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - FAD LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-000i-0003900000-db784762434fac3d351f | 2012-08-31 | HMDB team, MONA | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - FAD 10V, Positive-QTOF | splash10-000i-0932110400-f05626487523221c34de | 2016-09-12 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - FAD 20V, Positive-QTOF | splash10-000i-0930000000-86c3d6490bc3a27e053a | 2016-09-12 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - FAD 40V, Positive-QTOF | splash10-052r-0980000000-58a46c839e0e520474d7 | 2016-09-12 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - FAD 10V, Negative-QTOF | splash10-0006-8482400900-2c4839849821d2807c57 | 2016-09-12 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - FAD 20V, Negative-QTOF | splash10-001l-5920100000-b38889aa3d09576e2f3e | 2016-09-12 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - FAD 40V, Negative-QTOF | splash10-0a7l-3900000000-f7144f6355fab252a516 | 2016-09-12 | Wishart Lab | View Spectrum |
NMR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Experimental 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, H2O, experimental) | 2012-12-05 | Wishart Lab | View Spectrum |
|
|---|
| Biological Properties |
|---|
| Cellular Locations | - Cytoplasm
- Mitochondria
- Endoplasmic reticulum
- Peroxisome
|
|---|
| Biospecimen Locations | |
|---|
| Tissue Locations | - Erythrocyte
- Liver
- Placenta
- Skeletal Muscle
|
|---|
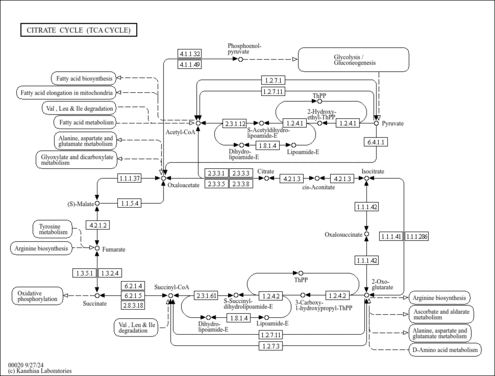
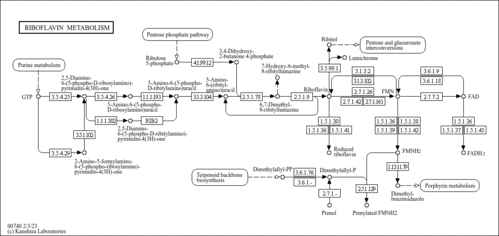
| Pathways | |
|---|
| Normal Concentrations |
|---|
| |
| Blood | Detected and Quantified | 0.078 +/- 0.054 uM | Adolescent (13-18 years old) | Female | Normal | | details | | Blood | Detected and Quantified | 0.075 (0.056-0.097) uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 0.061 (0.044-0.078) uM | Adult (>18 years old) | Female | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details |
|
|---|
| Abnormal Concentrations |
|---|
| |
| Blood | Detected and Quantified | 0.048 +/- 0.023 uM | Adolescent (13-18 years old) | Female | Anorexia nervosa | | details | | Blood | Detected and Quantified | 0.031 +/- 0.01 uM | Children (1-13 years old) | Both | Malnutrition (type kwashiorkor and marasmus) | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal Cancer | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details |
|
|---|
| Associated Disorders and Diseases |
|---|
| Disease References | | Anorexia nervosa |
|---|
- Capo-chichi CD, Gueant JL, Lefebvre E, Bennani N, Lorentz E, Vidailhet C, Vidailhet M: Riboflavin and riboflavin-derived cofactors in adolescent girls with anorexia nervosa. Am J Clin Nutr. 1999 Apr;69(4):672-8. [PubMed:10197568 ]
| | Colorectal cancer |
|---|
- Brown DG, Rao S, Weir TL, O'Malia J, Bazan M, Brown RJ, Ryan EP: Metabolomics and metabolic pathway networks from human colorectal cancers, adjacent mucosa, and stool. Cancer Metab. 2016 Jun 6;4:11. doi: 10.1186/s40170-016-0151-y. eCollection 2016. [PubMed:27275383 ]
- Goedert JJ, Sampson JN, Moore SC, Xiao Q, Xiong X, Hayes RB, Ahn J, Shi J, Sinha R: Fecal metabolomics: assay performance and association with colorectal cancer. Carcinogenesis. 2014 Sep;35(9):2089-96. doi: 10.1093/carcin/bgu131. Epub 2014 Jul 18. [PubMed:25037050 ]
|
|
|---|
| Associated OMIM IDs | |
|---|
| External Links |
|---|
| DrugBank ID | DB03147 |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FooDB ID | FDB022511 |
|---|
| KNApSAcK ID | C00001500 |
|---|
| Chemspider ID | 559059 |
|---|
| KEGG Compound ID | C00016 |
|---|
| BioCyc ID | FAD |
|---|
| BiGG ID | 33521 |
|---|
| Wikipedia Link | Flavin_adenine_dinucleotide |
|---|
| METLIN ID | 6106 |
|---|
| PubChem Compound | 643975 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 16238 |
|---|
| Food Biomarker Ontology | Not Available |
|---|
| VMH ID | FAD |
|---|
| MarkerDB ID | Not Available |
|---|
| Good Scents ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| Material Safety Data Sheet (MSDS) | Download (PDF) |
|---|
| General References | - Flatz G, Simmersbach F: Flavin adenine dinucleotide concentration in erythrocytes with normal and deficient glucose-6-phosphate dehydrogenase. Klin Wochenschr. 1970 Sep 1;48(17):1071-2. [PubMed:5523465 ]
- Zempleni J: Determination of riboflavin and flavocoenzymes in human blood plasma by high-performance liquid chromatography. Ann Nutr Metab. 1995;39(4):224-6. [PubMed:8546438 ]
- Becker K, Wilkinson AR: Flavin adenine dinucleotide levels in erythrocytes of very low birthweight infants under vitamin supplementation. Biol Neonate. 1993;63(2):80-5. [PubMed:8448258 ]
- Lisowsky T, Lee JE, Polimeno L, Francavilla A, Hofhaus G: Mammalian augmenter of liver regeneration protein is a sulfhydryl oxidase. Dig Liver Dis. 2001 Mar;33(2):173-80. [PubMed:11346147 ]
- Cimino JA, Jhangiani S, Schwartz E, Cooperman JM: Riboflavin metabolism in the hypothyroid human adult. Proc Soc Exp Biol Med. 1987 Feb;184(2):151-3. [PubMed:3809170 ]
- Kodentsova VM, Vrzhesinskaia OA, Alekseeva IA, Spirichev VB: [Comparison of biochemical criteria for supplying the human body with riboflavin]. Vopr Med Khim. 1991 Sep-Oct;37(5):76-9. [PubMed:1759408 ]
- Lopez-Anaya A, Mayersohn M: Quantification of riboflavin, riboflavin 5'-phosphate and flavin adenine dinucleotide in plasma and urine by high-performance liquid chromatography. J Chromatogr. 1987 Dec 25;423:105-13. [PubMed:3443641 ]
- Van Binsbergen CJ, Odink J, Van den Berg H, Koppeschaar H, Coelingh Bennink HJ: Nutritional status in anorexia nervosa: clinical chemistry, vitamins, iron and zinc. Eur J Clin Nutr. 1988 Nov;42(11):929-37. [PubMed:3074921 ]
- Mohrenweiser HW, Novotny JE: ACP1GUA-1--a low-activity variant of human erythrocyte acid phosphatase: association with increased glutathione reductase activity. Am J Hum Genet. 1982 May;34(3):425-33. [PubMed:7081221 ]
- Gianazza E, Vergani L, Wait R, Brizio C, Brambilla D, Begum S, Giancaspero TA, Conserva F, Eberini I, Bufano D, Angelini C, Pegoraro E, Tramontano A, Barile M: Coordinated and reversible reduction of enzymes involved in terminal oxidative metabolism in skeletal muscle mitochondria from a riboflavin-responsive, multiple acyl-CoA dehydrogenase deficiency patient. Electrophoresis. 2006 Mar;27(5-6):1182-98. [PubMed:16470778 ]
- Cimino JA, Noto RA, Fusco CL, Cooperman JM: Riboflavin metabolism in the hypothyroid newborn. Am J Clin Nutr. 1988 Mar;47(3):481-3. [PubMed:3348160 ]
- Elshenawy S, Pinney SE, Stuart T, Doulias PT, Zura G, Parry S, Elovitz MA, Bennett MJ, Bansal A, Strauss JF 3rd, Ischiropoulos H, Simmons RA: The Metabolomic Signature of the Placenta in Spontaneous Preterm Birth. Int J Mol Sci. 2020 Feb 4;21(3). pii: ijms21031043. doi: 10.3390/ijms21031043. [PubMed:32033212 ]
|
|---|