| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Detected and Quantified |
|---|
| Creation Date | 2005-11-16 15:48:42 UTC |
|---|
| Update Date | 2025-05-29 18:10:01 UTC |
|---|
| HMDB ID | HMDB0000172 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
| Common Name | Isoleucine |
|---|
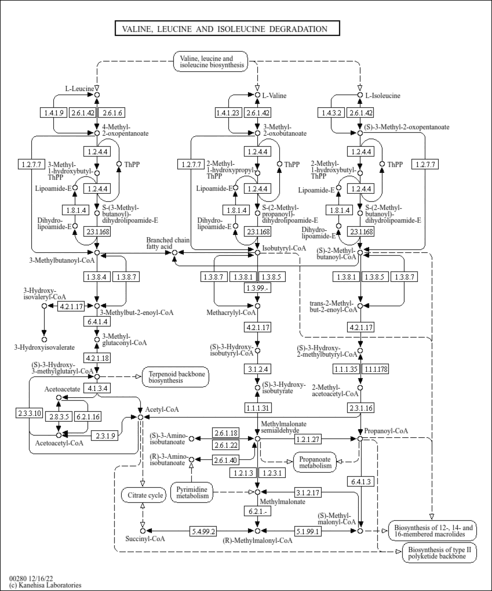
| Description | Isoleucine (Ile) or L-isoleucine is an alpha-amino acid. These are amino acids in which the amino group is attached to the carbon atom immediately adjacent to the carboxylate group (alpha carbon). Amino acids are organic compounds that contain amino (-NH2) and carboxyl (-COOH) functional groups, along with a side chain (R group) specific to each amino acid. L-isolecuine is one of 20 proteinogenic amino acids, i.e., the amino acids used in the biosynthesis of proteins. Isoleucine is found in all organisms ranging from bacteria to plants to animals. It is classified as a non-polar, uncharged (at physiological pH) aliphatic amino acid. Isoleucine is an essential amino acid in humans, meaning the body cannot synthesize it and that it must be obtained from the diet. In plants and microorganisms, isoleucine is synthesized starting from pyruvate and alpha-ketobutyrate. Isoleucine is classified as a branched chain amino acid (BCAA). BCAAs include three amino acids: isoleucine, leucine and valine. They are alpha amino acids whose carbon structure is marked by a beta branch point. Despite their structural similarities, BCAAs have different metabolic routes, with valine going solely to carbohydrates (glucogenic), leucine solely to fats (ketogenic) and isoleucine being both a glucogenic and a ketogenic amino acid. Isoleucine is catabolized via with alpha-ketoglutarate where upon it is oxidized and split into propionyl-CoA and acetyl-CoA. Propionyl-CoA is converted into succinyl-CoA, a TCA cycle intermediate which can be converted into oxaloacetate for gluconeogenesis (hence glucogenic). The acetyl-CoA can be fed into the TCA cycle by condensing with oxaloacetate to form citrate or used in the synthesis of ketone bodies or fatty acids. The different metabolism of BCAAs accounts for different requirements for these essential amino acids in humans: 12 mg/kg, 14 mg/kg and 16 mg/kg of valine, leucine and isoleucine are required respectively. Furthermore, these amino acids have different deficiency symptoms. Valine deficiency is marked by neurological defects in the brain, while isoleucine deficiency is marked by muscle tremors. BCAAs are decreased in patients with liver disease, such as hepatitis, hepatic coma, cirrhosis, extrahepatic biliary atresia. An inability to break down isoleucine, along with other amino acids, is associated with maple syrup urine disease (MSUD) (PMID: 34125801 ). Isoleucine, like other BCAAs, is associated with insulin resistance. In particular, higher levels of isoleucine are observed in the blood of diabetic mice, rats, and humans (PMID 25287287 ). Mice fed an isoleucine deprivation diet for one day have improved insulin sensitivity, and feeding of an isoleucine deprivation diet for one week significantly decreases blood glucose levels (PMID: 24684822 ). |
|---|
| Structure | InChI=1S/C6H13NO2/c1-3-4(2)5(7)6(8)9/h4-5H,3,7H2,1-2H3,(H,8,9)/t4-,5-/m0/s1 |
|---|
| Synonyms | | Value | Source |
|---|
| (2S,3S)-2-Amino-3-methylpentanoic acid | ChEBI | | 2-Amino-3-methylvaleric acid | ChEBI | | alpha-Amino-beta-methylvaleric acid | ChEBI | | I | ChEBI | | Ile | ChEBI | | ISOLEUCINE | ChEBI | | (2S,3S)-2-Amino-3-methylpentanoate | Generator | | 2-Amino-3-methylvalerate | Generator | | a-Amino-b-methylvalerate | Generator | | a-Amino-b-methylvaleric acid | Generator | | alpha-Amino-beta-methylvalerate | Generator | | Α-amino-β-methylvalerate | Generator | | Α-amino-β-methylvaleric acid | Generator | | (2S,3S)-2-Amino-3-methyl-pentanoate | HMDB | | (2S,3S)-2-Amino-3-methyl-pentanoic acid | HMDB | | (2S,3S)-a-Amino-b-methyl-N-valerate | HMDB | | (2S,3S)-a-Amino-b-methyl-N-valeric acid | HMDB | | (2S,3S)-a-Amino-b-methylvalerate | HMDB | | (2S,3S)-a-Amino-b-methylvaleric acid | HMDB | | (2S,3S)-Alph-amino-beta-methylvalerate | HMDB | | (2S,3S)-Alph-amino-beta-methylvaleric acid | HMDB | | (2S,3S)-alpha-Amino-beta-merthyl-N-valerate | HMDB | | (2S,3S)-alpha-Amino-beta-merthyl-N-valeric acid | HMDB | | (2S,3S)-alpha-Amino-beta-merthylvalerate | HMDB | | (2S,3S)-alpha-Amino-beta-merthylvaleric acid | HMDB | | (2S,3S)-alpha-Amino-beta-methyl-N-valerate | HMDB | | (2S,3S)-alpha-Amino-beta-methyl-N-valeric acid | HMDB | | (2S,3S)-alpha-Amino-beta-methylvalerate | HMDB | | (2S,3S)-alpha-Amino-beta-methylvaleric acid | HMDB | | (S)-Isoleucine | HMDB | | (S,S)-Isoleucine | HMDB | | 2-Amino-3-methylpentanoate | HMDB | | 2-Amino-3-methylpentanoic acid | HMDB | | 2S,3S-Isoleucine | HMDB | | Erythro-L-isoleucine | HMDB | | Iso-leucine | HMDB | | L-(+)-Isoleucine | HMDB | | L-Ile | HMDB | | [S-(R*,r*)]-2-amino-3-methylpentanoate | HMDB | | [S-(R*,r*)]-2-amino-3-methylpentanoic acid | HMDB | | Isoleucine, L-isomer | HMDB | | Alloisoleucine | HMDB | | Isoleucine, L isomer | HMDB | | L-Isomer isoleucine | HMDB | | 2S-Amino-3S-methylpentanoate | HMDB | | L-Isoleucine | KEGG |
|
|---|
| Chemical Formula | C6H13NO2 |
|---|
| Average Molecular Weight | 131.1729 |
|---|
| Monoisotopic Molecular Weight | 131.094628665 |
|---|
| IUPAC Name | (2S,3S)-2-amino-3-methylpentanoic acid |
|---|
| Traditional Name | L-isoleucine |
|---|
| CAS Registry Number | 73-32-5 |
|---|
| SMILES | CC[C@H](C)[C@H](N)C(O)=O |
|---|
| InChI Identifier | InChI=1S/C6H13NO2/c1-3-4(2)5(7)6(8)9/h4-5H,3,7H2,1-2H3,(H,8,9)/t4-,5-/m0/s1 |
|---|
| InChI Key | AGPKZVBTJJNPAG-WHFBIAKZSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as isoleucine and derivatives. Isoleucine and derivatives are compounds containing isoleucine or a derivative thereof resulting from reaction of isoleucine at the amino group or the carboxy group, or from the replacement of any hydrogen of glycine by a heteroatom. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Carboxylic acids and derivatives |
|---|
| Sub Class | Amino acids, peptides, and analogues |
|---|
| Direct Parent | Isoleucine and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Isoleucine or derivatives
- Alpha-amino acid
- L-alpha-amino acid
- Branched fatty acid
- Methyl-branched fatty acid
- Fatty acid
- Fatty acyl
- Amino acid
- Monocarboxylic acid or derivatives
- Carboxylic acid
- Organic oxide
- Organopnictogen compound
- Primary amine
- Organooxygen compound
- Organonitrogen compound
- Primary aliphatic amine
- Carbonyl group
- Organic oxygen compound
- Amine
- Organic nitrogen compound
- Hydrocarbon derivative
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Not Available | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | 285.5 °C | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | 35 mg/mL | Human Metabolome Project | | LogP | -1.70 | HANSCH,C ET AL. (1995) |
|
|---|
| Experimental Chromatographic Properties | Experimental Collision Cross Sections |
|---|
| Predicted Molecular Properties | |
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Retention Times Underivatized| Chromatographic Method | Retention Time | Reference |
|---|
| Measured using a Waters Acquity ultraperformance liquid chromatography (UPLC) ethylene-bridged hybrid (BEH) C18 column (100 mm × 2.1 mm; 1.7 μmparticle diameter). Predicted by Afia on May 17, 2022. Predicted by Afia on May 17, 2022. | 1.25 minutes | 32390414 | | Predicted by Siyang on May 30, 2022 | 9.9376 minutes | 33406817 | | Predicted by Siyang using ReTip algorithm on June 8, 2022 | 6.56 minutes | 32390414 | | AjsUoB = Accucore 150 Amide HILIC with 10mM Ammonium Formate, 0.1% Formic Acid | 317.8 seconds | 40023050 | | Fem_Long = Waters ACQUITY UPLC HSS T3 C18 with Water:MeOH and 0.1% Formic Acid | 692.6 seconds | 40023050 | | Fem_Lipids = Ascentis Express C18 with (60:40 water:ACN):(90:10 IPA:ACN) and 10mM NH4COOH + 0.1% Formic Acid | 326.9 seconds | 40023050 | | Life_Old = Waters ACQUITY UPLC BEH C18 with Water:(20:80 acetone:ACN) and 0.1% Formic Acid | 74.6 seconds | 40023050 | | Life_New = RP Waters ACQUITY UPLC HSS T3 C18 with Water:(30:70 MeOH:ACN) and 0.1% Formic Acid | 181.5 seconds | 40023050 | | RIKEN = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 59.7 seconds | 40023050 | | Eawag_XBridgeC18 = XBridge C18 3.5u 2.1x50 mm with Water:MeOH and 0.1% Formic Acid | 305.5 seconds | 40023050 | | BfG_NTS_RP1 =Agilent Zorbax Eclipse Plus C18 (2.1 mm x 150 mm, 3.5 um) with Water:ACN and 0.1% Formic Acid | 277.3 seconds | 40023050 | | HILIC_BDD_2 = Merck SeQuant ZIC-HILIC with ACN(0.1% formic acid):water(16 mM ammonium formate) | 549.9 seconds | 40023050 | | UniToyama_Atlantis = RP Waters Atlantis T3 (2.1 x 150 mm, 5 um) with ACN:Water and 0.1% Formic Acid | 663.4 seconds | 40023050 | | BDD_C18 = Hypersil Gold 1.9µm C18 with Water:ACN and 0.1% Formic Acid | 149.4 seconds | 40023050 | | UFZ_Phenomenex = Kinetex Core-Shell C18 2.6 um, 3.0 x 100 mm, Phenomenex with Water:MeOH and 0.1% Formic Acid | 778.9 seconds | 40023050 | | SNU_RIKEN_POS = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 180.2 seconds | 40023050 | | RPMMFDA = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 238.5 seconds | 40023050 | | MTBLS87 = Merck SeQuant ZIC-pHILIC column with ACN:Water and :ammonium carbonate | 490.4 seconds | 40023050 | | KI_GIAR_zic_HILIC_pH2_7 = Merck SeQuant ZIC-HILIC with ACN:Water and 0.1% FA | 393.3 seconds | 40023050 | | Meister zic-pHILIC pH9.3 = Merck SeQuant ZIC-pHILIC column with ACN:Water 5mM NH4Ac pH9.3 and 5mM ammonium acetate in water | 252.5 seconds | 40023050 |
Predicted Kovats Retention IndicesUnderivatizedDerivatized| Derivative Name / Structure | SMILES | Kovats RI Value | Column Type | Reference |
|---|
| L-Isoleucine,1TMS,isomer #1 | CC[C@H](C)[C@H](N)C(=O)O[Si](C)(C)C | 1180.4 | Semi standard non polar | 33892256 | | L-Isoleucine,1TMS,isomer #2 | CC[C@H](C)[C@H](N[Si](C)(C)C)C(=O)O | 1292.1 | Semi standard non polar | 33892256 | | L-Isoleucine,2TMS,isomer #1 | CC[C@H](C)[C@H](N[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1316.6 | Semi standard non polar | 33892256 | | L-Isoleucine,2TMS,isomer #1 | CC[C@H](C)[C@H](N[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1308.4 | Standard non polar | 33892256 | | L-Isoleucine,2TMS,isomer #1 | CC[C@H](C)[C@H](N[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1419.5 | Standard polar | 33892256 | | L-Isoleucine,2TMS,isomer #2 | CC[C@H](C)[C@@H](C(=O)O)N([Si](C)(C)C)[Si](C)(C)C | 1480.8 | Semi standard non polar | 33892256 | | L-Isoleucine,2TMS,isomer #2 | CC[C@H](C)[C@@H](C(=O)O)N([Si](C)(C)C)[Si](C)(C)C | 1322.9 | Standard non polar | 33892256 | | L-Isoleucine,2TMS,isomer #2 | CC[C@H](C)[C@@H](C(=O)O)N([Si](C)(C)C)[Si](C)(C)C | 1511.7 | Standard polar | 33892256 | | L-Isoleucine,3TMS,isomer #1 | CC[C@H](C)[C@@H](C(=O)O[Si](C)(C)C)N([Si](C)(C)C)[Si](C)(C)C | 1501.3 | Semi standard non polar | 33892256 | | L-Isoleucine,3TMS,isomer #1 | CC[C@H](C)[C@@H](C(=O)O[Si](C)(C)C)N([Si](C)(C)C)[Si](C)(C)C | 1434.9 | Standard non polar | 33892256 | | L-Isoleucine,3TMS,isomer #1 | CC[C@H](C)[C@@H](C(=O)O[Si](C)(C)C)N([Si](C)(C)C)[Si](C)(C)C | 1401.2 | Standard polar | 33892256 | | L-Isoleucine,1TBDMS,isomer #1 | CC[C@H](C)[C@H](N)C(=O)O[Si](C)(C)C(C)(C)C | 1396.9 | Semi standard non polar | 33892256 | | L-Isoleucine,1TBDMS,isomer #2 | CC[C@H](C)[C@H](N[Si](C)(C)C(C)(C)C)C(=O)O | 1519.3 | Semi standard non polar | 33892256 | | L-Isoleucine,2TBDMS,isomer #1 | CC[C@H](C)[C@H](N[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 1749.9 | Semi standard non polar | 33892256 | | L-Isoleucine,2TBDMS,isomer #1 | CC[C@H](C)[C@H](N[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 1724.7 | Standard non polar | 33892256 | | L-Isoleucine,2TBDMS,isomer #1 | CC[C@H](C)[C@H](N[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 1722.9 | Standard polar | 33892256 | | L-Isoleucine,2TBDMS,isomer #2 | CC[C@H](C)[C@@H](C(=O)O)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 1878.2 | Semi standard non polar | 33892256 | | L-Isoleucine,2TBDMS,isomer #2 | CC[C@H](C)[C@@H](C(=O)O)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 1758.8 | Standard non polar | 33892256 | | L-Isoleucine,2TBDMS,isomer #2 | CC[C@H](C)[C@@H](C(=O)O)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 1750.4 | Standard polar | 33892256 | | L-Isoleucine,3TBDMS,isomer #1 | CC[C@H](C)[C@@H](C(=O)O[Si](C)(C)C(C)(C)C)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2161.4 | Semi standard non polar | 33892256 | | L-Isoleucine,3TBDMS,isomer #1 | CC[C@H](C)[C@@H](C(=O)O[Si](C)(C)C(C)(C)C)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2055.1 | Standard non polar | 33892256 | | L-Isoleucine,3TBDMS,isomer #1 | CC[C@H](C)[C@@H](C(=O)O[Si](C)(C)C(C)(C)C)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 1828.2 | Standard polar | 33892256 |
|
|---|
| GC-MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental GC-MS | GC-MS Spectrum - Isoleucine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (2 TMS) | splash10-0a4i-0930000000-e78f845bb2a8d4736476 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Isoleucine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (2 TMS) | splash10-0a4i-0910000000-de9162d149073d0e2a37 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Isoleucine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (2 TMS) | splash10-0a4i-0920000000-599e61f8ccdb6525c7a9 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Isoleucine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (Non-derivatized) | splash10-0a4i-0910000000-742e44c426c0d6c2a9ac | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Isoleucine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (2 TMS) | splash10-05fr-8910000000-dbb33e0f02ac2ca5fedd | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Isoleucine EI-B (Non-derivatized) | splash10-0a4i-0920000000-37690f426455ac41e2c2 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Isoleucine GC-EI-TOF (Non-derivatized) | splash10-0a4i-0930000000-e78f845bb2a8d4736476 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Isoleucine GC-EI-TOF (Non-derivatized) | splash10-0a4i-0910000000-de9162d149073d0e2a37 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Isoleucine GC-EI-TOF (Non-derivatized) | splash10-0a4i-0920000000-599e61f8ccdb6525c7a9 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Isoleucine GC-EI-TOF (Non-derivatized) | splash10-0a4i-0910000000-742e44c426c0d6c2a9ac | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Isoleucine GC-EI-QQ (Non-derivatized) | splash10-0udi-2392000000-74cca1eb265b8e3d7f43 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Isoleucine GC-EI-TOF (Non-derivatized) | splash10-05fr-8910000000-dbb33e0f02ac2ca5fedd | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Isoleucine GC-EI-TOF (Non-derivatized) | splash10-000i-9100000000-27891dff695c5db53796 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Isoleucine GC-EI-TOF (Non-derivatized) | splash10-0a4i-0910000000-43ec1890f6dbd6aea657 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Isoleucine GC-MS (Non-derivatized) - 70eV, Positive | splash10-00b9-9100000000-40bc9ef43da5f18be883 | 2016-09-22 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Isoleucine GC-MS (1 TMS) - 70eV, Positive | splash10-000i-9200000000-ae8694c0f3e4dcc608f9 | 2017-10-06 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Isoleucine GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Isoleucine GC-MS (TMS_1_2) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Isoleucine GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Isoleucine GC-MS (TBDMS_1_2) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-004r-9000000000-34d4d4cb7042da231eb4 | 2014-09-20 | Not Available | View Spectrum |
MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine Quattro_QQQ 10V, Positive-QTOF (Annotated) | splash10-000i-9300000000-8a23b3e62231eb65f80f | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine Quattro_QQQ 25V, Positive-QTOF (Annotated) | splash10-00ko-9000000000-ac7e71578d7e1d0e8ce9 | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine Quattro_QQQ 40V, Positive-QTOF (Annotated) | splash10-052f-9000000000-2586d4a089921dd977ed | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-001i-0900000000-5b11521ff6a631376d2b | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-000i-9000000000-3c352f229e4067fcd489 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-000i-9000000000-8a7ef48fc0c1b6f845c2 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-0002-0920000000-2e11aa1c7a5defc386db | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-001i-0900000000-b63ac9cf06eda4c54a81 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-000i-9000000000-e5b0c8c6b09f541d6dbf | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-000i-9000000000-d89d16d5c2242e44ec63 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-000i-9000000000-5de583142c7bdb381c8f | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine LC-ESI-QQ (API3000, Applied Biosystems) 10V, Negative-QTOF | splash10-001i-0900000000-8c75bde35c4a4073ee65 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine LC-ESI-QQ (API3000, Applied Biosystems) 20V, Negative-QTOF | splash10-001i-0900000000-233a9862f616afea2d17 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine LC-ESI-QQ (API3000, Applied Biosystems) 30V, Negative-QTOF | splash10-001i-1900000000-375f35065b82b13e6d6e | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine LC-ESI-QQ (API3000, Applied Biosystems) 40V, Negative-QTOF | splash10-00dl-9000000000-d0d4a9f90fe2aca74483 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine LC-ESI-QQ (API3000, Applied Biosystems) 50V, Negative-QTOF | splash10-0006-9000000000-0018f47571feaf232ff8 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine LC-ESI-QQ (API3000, Applied Biosystems) 10V, Positive-QTOF | splash10-001r-7900000000-278dc67396114be331ba | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine LC-ESI-QQ (API3000, Applied Biosystems) 20V, Positive-QTOF | splash10-000i-9000000000-e26c042aa6231eeca071 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine LC-ESI-QQ (API3000, Applied Biosystems) 30V, Positive-QTOF | splash10-014r-9000000000-b6c1752fd3fbccb3d1c5 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine LC-ESI-QQ (API3000, Applied Biosystems) 40V, Positive-QTOF | splash10-05mo-9000000000-0569c3162621252ed0a8 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine LC-ESI-QQ (API3000, Applied Biosystems) 50V, Positive-QTOF | splash10-052f-9000000000-3d52b5d56d3fab45276e | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Isoleucine CE-ESI-TOF (CE-system connected to 6210 Time-of-Flight MS, Agilent) , Positive-QTOF | splash10-001i-0900000000-720554d58264a9cfdb67 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Isoleucine 10V, Positive-QTOF | splash10-0019-9500000000-00268b2694cde281641b | 2016-09-12 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Isoleucine 20V, Positive-QTOF | splash10-000i-9000000000-4014793e17e0290790a4 | 2016-09-12 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Isoleucine 40V, Positive-QTOF | splash10-0a4i-9000000000-eb9a97a14461e524e76e | 2016-09-12 | Wishart Lab | View Spectrum |
NMR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Experimental 1D NMR | 1H NMR Spectrum (1D, 600 MHz, H2O, experimental) | 2012-12-04 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Experimental 1D NMR | 13C NMR Spectrum (1D, 400 MHz, H2O, experimental) | 2021-10-10 | Wishart Lab | View Spectrum | | Experimental 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, H2O, experimental) | 2012-12-05 | Wishart Lab | View Spectrum |
IR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted IR Spectrum | IR Ion Spectrum (Quadrupole Ion Trap, ESI+, Adduct: [M+H]+) | 2022-02-11 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M-H]-) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+H]+) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+Na]+) | 2023-02-03 | FELIX lab | View Spectrum |
|
|---|
| Disease References | | Alzheimer's disease |
|---|
- Fonteh AN, Harrington RJ, Tsai A, Liao P, Harrington MG: Free amino acid and dipeptide changes in the body fluids from Alzheimer's disease subjects. Amino Acids. 2007 Feb;32(2):213-24. Epub 2006 Oct 10. [PubMed:17031479 ]
- Tsuruoka M, Hara J, Hirayama A, Sugimoto M, Soga T, Shankle WR, Tomita M: Capillary electrophoresis-mass spectrometry-based metabolome analysis of serum and saliva from neurodegenerative dementia patients. Electrophoresis. 2013 Oct;34(19):2865-72. doi: 10.1002/elps.201300019. Epub 2013 Sep 6. [PubMed:23857558 ]
| | Heart failure |
|---|
- Norrelund H, Wiggers H, Halbirk M, Frystyk J, Flyvbjerg A, Botker HE, Schmitz O, Jorgensen JO, Christiansen JS, Moller N: Abnormalities of whole body protein turnover, muscle metabolism and levels of metabolic hormones in patients with chronic heart failure. J Intern Med. 2006 Jul;260(1):11-21. [PubMed:16789974 ]
| | Maple syrup urine disease |
|---|
- Shigematsu Y, Kikuchi K, Momoi T, Sudo M, Kikawa Y, Nosaka K, Kuriyama M, Haruki S, Sanada K, Hamano N, et al.: Organic acids and branched-chain amino acids in body fluids before and after multiple exchange transfusions in maple syrup urine disease. J Inherit Metab Dis. 1983;6(4):183-9. [PubMed:6422161 ]
- Yunus ZM, Kamaludin DA, Mamat M, Choy YS, Ngu L: Clinical and biochemical profiles of maple syrup urine disease in malaysian children. JIMD Rep. 2012;5:99-107. doi: 10.1007/8904_2011_105. Epub 2011 Dec 11. [PubMed:23430924 ]
- Barschak AG, Marchesan C, Sitta A, Deon M, Giugliani R, Wajner M, Vargas CR: Maple syrup urine disease in treated patients: biochemical and oxidative stress profiles. Clin Biochem. 2008 Mar;41(4-5):317-24. Epub 2007 Dec 5. [PubMed:18088602 ]
- G.Frauendienst-Egger, Friedrich K. Trefz (2017). MetaGene: Metabolic & Genetic Information Center (MIC: http://www.metagene.de). METAGENE consortium.
- Wendel, U., Becker, K., Przyrembel, H. et al. (1980). Peritoneal dialysis in maple-syrup-urine disease: Studies on branched-chain amino and keto acids. Eur J Pediatr (1980) 134: 57. https://doi.org/10.1007/BF00442404. Eur J Pediatr.
| | Lipoyltransferase 1 Deficiency |
|---|
- Soreze Y, Boutron A, Habarou F, Barnerias C, Nonnenmacher L, Delpech H, Mamoune A, Chretien D, Hubert L, Bole-Feysot C, Nitschke P, Correia I, Sardet C, Boddaert N, Hamel Y, Delahodde A, Ottolenghi C, de Lonlay P: Mutations in human lipoyltransferase gene LIPT1 cause a Leigh disease with secondary deficiency for pyruvate and alpha-ketoglutarate dehydrogenase. Orphanet J Rare Dis. 2013 Dec 17;8:192. doi: 10.1186/1750-1172-8-192. [PubMed:24341803 ]
| | Branched-chain Keto Acid Dehydrogenase Kinase Deficiency |
|---|
- Novarino G, El-Fishawy P, Kayserili H, Meguid NA, Scott EM, Schroth J, Silhavy JL, Kara M, Khalil RO, Ben-Omran T, Ercan-Sencicek AG, Hashish AF, Sanders SJ, Gupta AR, Hashem HS, Matern D, Gabriel S, Sweetman L, Rahimi Y, Harris RA, State MW, Gleeson JG: Mutations in BCKD-kinase lead to a potentially treatable form of autism with epilepsy. Science. 2012 Oct 19;338(6105):394-7. doi: 10.1126/science.1224631. Epub 2012 Sep 6. [PubMed:22956686 ]
| | Dihydrolipoamide Dehydrogenase Deficiency |
|---|
- Kuhara T, Shinka T, Inoue Y, Matsumoto M, Yoshino M, Sakaguchi Y, Matsumoto I: Studies of urinary organic acid profiles of a patient with dihydrolipoyl dehydrogenase deficiency. Clin Chim Acta. 1983 Sep 30;133(2):133-40. [PubMed:6688766 ]
| | Saccharopinuria |
|---|
- Simell O, Visakorpi JK, Donner M: Saccharopinuria. Arch Dis Child. 1972 Feb;47(251):52-5. [PubMed:5018656 ]
| | Pregnancy |
|---|
- Bahado-Singh RO, Akolekar R, Mandal R, Dong E, Xia J, Kruger M, Wishart DS, Nicolaides K: Metabolomics and first-trimester prediction of early-onset preeclampsia. J Matern Fetal Neonatal Med. 2012 Oct;25(10):1840-7. doi: 10.3109/14767058.2012.680254. Epub 2012 Apr 28. [PubMed:22494326 ]
- Bahado-Singh RO, Akolekar R, Chelliah A, Mandal R, Dong E, Kruger M, Wishart DS, Nicolaides K: Metabolomic analysis for first-trimester trisomy 18 detection. Am J Obstet Gynecol. 2013 Jul;209(1):65.e1-9. doi: 10.1016/j.ajog.2013.03.028. Epub 2013 Mar 25. [PubMed:23535240 ]
- Bahado-Singh RO, Ertl R, Mandal R, Bjorndahl TC, Syngelaki A, Han B, Dong E, Liu PB, Alpay-Savasan Z, Wishart DS, Nicolaides KH: Metabolomic prediction of fetal congenital heart defect in the first trimester. Am J Obstet Gynecol. 2014 Sep;211(3):240.e1-240.e14. doi: 10.1016/j.ajog.2014.03.056. Epub 2014 Apr 1. [PubMed:24704061 ]
| | Early preeclampsia |
|---|
- Bahado-Singh RO, Akolekar R, Mandal R, Dong E, Xia J, Kruger M, Wishart DS, Nicolaides K: Metabolomics and first-trimester prediction of early-onset preeclampsia. J Matern Fetal Neonatal Med. 2012 Oct;25(10):1840-7. doi: 10.3109/14767058.2012.680254. Epub 2012 Apr 28. [PubMed:22494326 ]
| | Epilepsy |
|---|
- Rainesalo S, Keranen T, Palmio J, Peltola J, Oja SS, Saransaari P: Plasma and cerebrospinal fluid amino acids in epileptic patients. Neurochem Res. 2004 Jan;29(1):319-24. [PubMed:14992292 ]
| | Colorectal cancer |
|---|
- Ritchie SA, Ahiahonu PW, Jayasinghe D, Heath D, Liu J, Lu Y, Jin W, Kavianpour A, Yamazaki Y, Khan AM, Hossain M, Su-Myat KK, Wood PL, Krenitsky K, Takemasa I, Miyake M, Sekimoto M, Monden M, Matsubara H, Nomura F, Goodenowe DB: Reduced levels of hydroxylated, polyunsaturated ultra long-chain fatty acids in the serum of colorectal cancer patients: implications for early screening and detection. BMC Med. 2010 Feb 15;8:13. doi: 10.1186/1741-7015-8-13. [PubMed:20156336 ]
- Ni Y, Xie G, Jia W: Metabonomics of human colorectal cancer: new approaches for early diagnosis and biomarker discovery. J Proteome Res. 2014 Sep 5;13(9):3857-70. doi: 10.1021/pr500443c. Epub 2014 Aug 14. [PubMed:25105552 ]
- Lin Y, Ma C, Liu C, Wang Z, Yang J, Liu X, Shen Z, Wu R: NMR-based fecal metabolomics fingerprinting as predictors of earlier diagnosis in patients with colorectal cancer. Oncotarget. 2016 May 17;7(20):29454-64. doi: 10.18632/oncotarget.8762. [PubMed:27107423 ]
- Brown DG, Rao S, Weir TL, O'Malia J, Bazan M, Brown RJ, Ryan EP: Metabolomics and metabolic pathway networks from human colorectal cancers, adjacent mucosa, and stool. Cancer Metab. 2016 Jun 6;4:11. doi: 10.1186/s40170-016-0151-y. eCollection 2016. [PubMed:27275383 ]
- Sinha R, Ahn J, Sampson JN, Shi J, Yu G, Xiong X, Hayes RB, Goedert JJ: Fecal Microbiota, Fecal Metabolome, and Colorectal Cancer Interrelations. PLoS One. 2016 Mar 25;11(3):e0152126. doi: 10.1371/journal.pone.0152126. eCollection 2016. [PubMed:27015276 ]
- Goedert JJ, Sampson JN, Moore SC, Xiao Q, Xiong X, Hayes RB, Ahn J, Shi J, Sinha R: Fecal metabolomics: assay performance and association with colorectal cancer. Carcinogenesis. 2014 Sep;35(9):2089-96. doi: 10.1093/carcin/bgu131. Epub 2014 Jul 18. [PubMed:25037050 ]
| | Leukemia |
|---|
- Peng CT, Wu KH, Lan SJ, Tsai JJ, Tsai FJ, Tsai CH: Amino acid concentrations in cerebrospinal fluid in children with acute lymphoblastic leukemia undergoing chemotherapy. Eur J Cancer. 2005 May;41(8):1158-63. Epub 2005 Apr 14. [PubMed:15911239 ]
| | Schizophrenia |
|---|
- Do KQ, Lauer CJ, Schreiber W, Zollinger M, Gutteck-Amsler U, Cuenod M, Holsboer F: gamma-Glutamylglutamine and taurine concentrations are decreased in the cerebrospinal fluid of drug-naive patients with schizophrenic disorders. J Neurochem. 1995 Dec;65(6):2652-62. [PubMed:7595563 ]
- Bjerkenstedt L, Edman G, Hagenfeldt L, Sedvall G, Wiesel FA: Plasma amino acids in relation to cerebrospinal fluid monoamine metabolites in schizophrenic patients and healthy controls. Br J Psychiatry. 1985 Sep;147:276-82. [PubMed:2415198 ]
- Yang J, Chen T, Sun L, Zhao Z, Qi X, Zhou K, Cao Y, Wang X, Qiu Y, Su M, Zhao A, Wang P, Yang P, Wu J, Feng G, He L, Jia W, Wan C: Potential metabolite markers of schizophrenia. Mol Psychiatry. 2013 Jan;18(1):67-78. doi: 10.1038/mp.2011.131. Epub 2011 Oct 25. [PubMed:22024767 ]
| | Ulcerative colitis |
|---|
- Marchesi JR, Holmes E, Khan F, Kochhar S, Scanlan P, Shanahan F, Wilson ID, Wang Y: Rapid and noninvasive metabonomic characterization of inflammatory bowel disease. J Proteome Res. 2007 Feb;6(2):546-51. [PubMed:17269711 ]
- Le Gall G, Noor SO, Ridgway K, Scovell L, Jamieson C, Johnson IT, Colquhoun IJ, Kemsley EK, Narbad A: Metabolomics of fecal extracts detects altered metabolic activity of gut microbiota in ulcerative colitis and irritable bowel syndrome. J Proteome Res. 2011 Sep 2;10(9):4208-18. doi: 10.1021/pr2003598. Epub 2011 Aug 8. [PubMed:21761941 ]
- Bjerrum JT, Wang Y, Hao F, Coskun M, Ludwig C, Gunther U, Nielsen OH: Metabonomics of human fecal extracts characterize ulcerative colitis, Crohn's disease and healthy individuals. Metabolomics. 2015;11:122-133. Epub 2014 Jun 1. [PubMed:25598765 ]
- Kolho KL, Pessia A, Jaakkola T, de Vos WM, Velagapudi V: Faecal and Serum Metabolomics in Paediatric Inflammatory Bowel Disease. J Crohns Colitis. 2017 Mar 1;11(3):321-334. doi: 10.1093/ecco-jcc/jjw158. [PubMed:27609529 ]
| | Autism |
|---|
- De Angelis M, Piccolo M, Vannini L, Siragusa S, De Giacomo A, Serrazzanetti DI, Cristofori F, Guerzoni ME, Gobbetti M, Francavilla R: Fecal microbiota and metabolome of children with autism and pervasive developmental disorder not otherwise specified. PLoS One. 2013 Oct 9;8(10):e76993. doi: 10.1371/journal.pone.0076993. eCollection 2013. [PubMed:24130822 ]
| | Irritable bowel syndrome |
|---|
- Le Gall G, Noor SO, Ridgway K, Scovell L, Jamieson C, Johnson IT, Colquhoun IJ, Kemsley EK, Narbad A: Metabolomics of fecal extracts detects altered metabolic activity of gut microbiota in ulcerative colitis and irritable bowel syndrome. J Proteome Res. 2011 Sep 2;10(9):4208-18. doi: 10.1021/pr2003598. Epub 2011 Aug 8. [PubMed:21761941 ]
- Hong YS, Hong KS, Park MH, Ahn YT, Lee JH, Huh CS, Lee J, Kim IK, Hwang GS, Kim JS: Metabonomic understanding of probiotic effects in humans with irritable bowel syndrome. J Clin Gastroenterol. 2011 May-Jun;45(5):415-25. doi: 10.1097/MCG.0b013e318207f76c. [PubMed:21494186 ]
| | Crohn's disease |
|---|
- Marchesi JR, Holmes E, Khan F, Kochhar S, Scanlan P, Shanahan F, Wilson ID, Wang Y: Rapid and noninvasive metabonomic characterization of inflammatory bowel disease. J Proteome Res. 2007 Feb;6(2):546-51. [PubMed:17269711 ]
- Bjerrum JT, Wang Y, Hao F, Coskun M, Ludwig C, Gunther U, Nielsen OH: Metabonomics of human fecal extracts characterize ulcerative colitis, Crohn's disease and healthy individuals. Metabolomics. 2015;11:122-133. Epub 2014 Jun 1. [PubMed:25598765 ]
- Kolho KL, Pessia A, Jaakkola T, de Vos WM, Velagapudi V: Faecal and Serum Metabolomics in Paediatric Inflammatory Bowel Disease. J Crohns Colitis. 2017 Mar 1;11(3):321-334. doi: 10.1093/ecco-jcc/jjw158. [PubMed:27609529 ]
| | Lewy body disease |
|---|
- Tsuruoka M, Hara J, Hirayama A, Sugimoto M, Soga T, Shankle WR, Tomita M: Capillary electrophoresis-mass spectrometry-based metabolome analysis of serum and saliva from neurodegenerative dementia patients. Electrophoresis. 2013 Oct;34(19):2865-72. doi: 10.1002/elps.201300019. Epub 2013 Sep 6. [PubMed:23857558 ]
| | Frontotemporal dementia |
|---|
- Tsuruoka M, Hara J, Hirayama A, Sugimoto M, Soga T, Shankle WR, Tomita M: Capillary electrophoresis-mass spectrometry-based metabolome analysis of serum and saliva from neurodegenerative dementia patients. Electrophoresis. 2013 Oct;34(19):2865-72. doi: 10.1002/elps.201300019. Epub 2013 Sep 6. [PubMed:23857558 ]
| | Periodontal disease |
|---|
- Sugimoto M, Wong DT, Hirayama A, Soga T, Tomita M: Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics. 2010 Mar;6(1):78-95. Epub 2009 Sep 10. [PubMed:20300169 ]
| | Pancreatic cancer |
|---|
- Sugimoto M, Wong DT, Hirayama A, Soga T, Tomita M: Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics. 2010 Mar;6(1):78-95. Epub 2009 Sep 10. [PubMed:20300169 ]
- OuYang D, Xu J, Huang H, Chen Z: Metabolomic profiling of serum from human pancreatic cancer patients using 1H NMR spectroscopy and principal component analysis. Appl Biochem Biotechnol. 2011 Sep;165(1):148-54. doi: 10.1007/s12010-011-9240-0. Epub 2011 Apr 20. [PubMed:21505807 ]
- Zhang L, Jin H, Guo X, Yang Z, Zhao L, Tang S, Mo P, Wu K, Nie Y, Pan Y, Fan D: Distinguishing pancreatic cancer from chronic pancreatitis and healthy individuals by (1)H nuclear magnetic resonance-based metabonomic profiles. Clin Biochem. 2012 Sep;45(13-14):1064-9. doi: 10.1016/j.clinbiochem.2012.05.012. Epub 2012 May 19. [PubMed:22613268 ]
| | Perillyl alcohol administration for cancer treatment |
|---|
- Sugimoto M, Wong DT, Hirayama A, Soga T, Tomita M: Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics. 2010 Mar;6(1):78-95. Epub 2009 Sep 10. [PubMed:20300169 ]
| | Attachment loss |
|---|
- Liebsch C, Pitchika V, Pink C, Samietz S, Kastenmuller G, Artati A, Suhre K, Adamski J, Nauck M, Volzke H, Friedrich N, Kocher T, Holtfreter B, Pietzner M: The Saliva Metabolome in Association to Oral Health Status. J Dent Res. 2019 Jun;98(6):642-651. doi: 10.1177/0022034519842853. Epub 2019 Apr 26. [PubMed:31026179 ]
| | Missing teeth |
|---|
- Liebsch C, Pitchika V, Pink C, Samietz S, Kastenmuller G, Artati A, Suhre K, Adamski J, Nauck M, Volzke H, Friedrich N, Kocher T, Holtfreter B, Pietzner M: The Saliva Metabolome in Association to Oral Health Status. J Dent Res. 2019 Jun;98(6):642-651. doi: 10.1177/0022034519842853. Epub 2019 Apr 26. [PubMed:31026179 ]
| | Periodontal Probing Depth |
|---|
- Liebsch C, Pitchika V, Pink C, Samietz S, Kastenmuller G, Artati A, Suhre K, Adamski J, Nauck M, Volzke H, Friedrich N, Kocher T, Holtfreter B, Pietzner M: The Saliva Metabolome in Association to Oral Health Status. J Dent Res. 2019 Jun;98(6):642-651. doi: 10.1177/0022034519842853. Epub 2019 Apr 26. [PubMed:31026179 ]
| | Eosinophilic esophagitis |
|---|
- Slae, M., Huynh, H., Wishart, D.S. (2014). Analysis of 30 normal pediatric urine samples via NMR spectroscopy (unpublished work). NA.
| | Autosomal dominant polycystic kidney disease |
|---|
- Gronwald W, Klein MS, Zeltner R, Schulze BD, Reinhold SW, Deutschmann M, Immervoll AK, Boger CA, Banas B, Eckardt KU, Oefner PJ: Detection of autosomal dominant polycystic kidney disease by NMR spectroscopic fingerprinting of urine. Kidney Int. 2011 Jun;79(11):1244-53. doi: 10.1038/ki.2011.30. Epub 2011 Mar 9. [PubMed:21389975 ]
|
|
|---|
| General References | - Sreekumar A, Poisson LM, Rajendiran TM, Khan AP, Cao Q, Yu J, Laxman B, Mehra R, Lonigro RJ, Li Y, Nyati MK, Ahsan A, Kalyana-Sundaram S, Han B, Cao X, Byun J, Omenn GS, Ghosh D, Pennathur S, Alexander DC, Berger A, Shuster JR, Wei JT, Varambally S, Beecher C, Chinnaiyan AM: Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature. 2009 Feb 12;457(7231):910-4. doi: 10.1038/nature07762. [PubMed:19212411 ]
- Silwood CJ, Lynch E, Claxson AW, Grootveld MC: 1H and (13)C NMR spectroscopic analysis of human saliva. J Dent Res. 2002 Jun;81(6):422-7. [PubMed:12097436 ]
- Nicholson JK, O'Flynn MP, Sadler PJ, Macleod AF, Juul SM, Sonksen PH: Proton-nuclear-magnetic-resonance studies of serum, plasma and urine from fasting normal and diabetic subjects. Biochem J. 1984 Jan 15;217(2):365-75. [PubMed:6696735 ]
- Engelborghs S, Marescau B, De Deyn PP: Amino acids and biogenic amines in cerebrospinal fluid of patients with Parkinson's disease. Neurochem Res. 2003 Aug;28(8):1145-50. [PubMed:12834252 ]
- Hagenfeldt L, Bjerkenstedt L, Edman G, Sedvall G, Wiesel FA: Amino acids in plasma and CSF and monoamine metabolites in CSF: interrelationship in healthy subjects. J Neurochem. 1984 Mar;42(3):833-7. [PubMed:6198473 ]
- Peng CT, Wu KH, Lan SJ, Tsai JJ, Tsai FJ, Tsai CH: Amino acid concentrations in cerebrospinal fluid in children with acute lymphoblastic leukemia undergoing chemotherapy. Eur J Cancer. 2005 May;41(8):1158-63. Epub 2005 Apr 14. [PubMed:15911239 ]
- Cynober LA: Plasma amino acid levels with a note on membrane transport: characteristics, regulation, and metabolic significance. Nutrition. 2002 Sep;18(9):761-6. [PubMed:12297216 ]
- Rainesalo S, Keranen T, Palmio J, Peltola J, Oja SS, Saransaari P: Plasma and cerebrospinal fluid amino acids in epileptic patients. Neurochem Res. 2004 Jan;29(1):319-24. [PubMed:14992292 ]
- He XY, Yang SY: Roles of type 10 17beta-hydroxysteroid dehydrogenase in intracrinology and metabolism of isoleucine and fatty acids. Endocr Metab Immune Disord Drug Targets. 2006 Mar;6(1):95-102. [PubMed:16611167 ]
- Vaalasti A, Suomalainen H, Kuokkanen K, Rechardt L: Neuropeptides in cutaneous neurofibromas of von Recklinghausen's disease. J Cutan Pathol. 1990 Dec;17(6):371-3. [PubMed:1981573 ]
- Eriste E, Norberg A, Bonetto V, Nepomuceno D, Lovenberg TW, Sillard R, Jornvall H: A C-terminally elongated form of PHI from porcine intestine. Cell Mol Life Sci. 1999 Nov 15;56(7-8):709-13. [PubMed:11212317 ]
- Jalan R, Olde Damink SW, Lui HF, Glabus M, Deutz NE, Hayes PC, Ebmeier K: Oral amino acid load mimicking hemoglobin results in reduced regional cerebral perfusion and deterioration in memory tests in patients with cirrhosis of the liver. Metab Brain Dis. 2003 Mar;18(1):37-49. [PubMed:12603081 ]
- Hervieu G, Segretain D, Nahon JL: Developmental and stage-dependent expression of melanin-concentrating hormone in mammalian germ cells. Biol Reprod. 1996 Jun;54(6):1161-72. [PubMed:8724342 ]
- Suk FM, Lin MH, Newman M, Pan S, Chen SH, Liu JD, Shih C: Replication advantage and host factor-independent phenotypes attributable to a common naturally occurring capsid mutation (I97L) in human hepatitis B virus. J Virol. 2002 Dec;76(23):12069-77. [PubMed:12414948 ]
- De Miranda J, Panizzutti R, Foltyn VN, Wolosker H: Cofactors of serine racemase that physiologically stimulate the synthesis of the N-methyl-D-aspartate (NMDA) receptor coagonist D-serine. Proc Natl Acad Sci U S A. 2002 Oct 29;99(22):14542-7. Epub 2002 Oct 22. [PubMed:12393813 ]
- Blomstrand E, Eliasson J, Karlsson HK, Kohnke R: Branched-chain amino acids activate key enzymes in protein synthesis after physical exercise. J Nutr. 2006 Jan;136(1 Suppl):269S-73S. [PubMed:16365096 ]
- Sato T, Shimada Y, Nagasawa N, Nakanishi S, Jingami H: Amino acid mutagenesis of the ligand binding site and the dimer interface of the metabotropic glutamate receptor 1. Identification of crucial residues for setting the activated state. J Biol Chem. 2003 Feb 7;278(6):4314-21. Epub 2002 Nov 19. [PubMed:12444084 ]
- Edvinsson L: Innervation and effects of dilatory neuropeptides on cerebral vessels. New aspects. Blood Vessels. 1991;28(1-3):35-45. [PubMed:2001478 ]
- Elshenawy S, Pinney SE, Stuart T, Doulias PT, Zura G, Parry S, Elovitz MA, Bennett MJ, Bansal A, Strauss JF 3rd, Ischiropoulos H, Simmons RA: The Metabolomic Signature of the Placenta in Spontaneous Preterm Birth. Int J Mol Sci. 2020 Feb 4;21(3). pii: ijms21031043. doi: 10.3390/ijms21031043. [PubMed:32033212 ]
- Allahwala A, Ahmed S, Afroze B: Maple syrup urine disease: magnetic resonance imaging findings in three patients. J Pak Med Assoc. 2021 Apr;71(4):1309-1313. doi: 10.47391/JPMA.1341. [PubMed:34125801 ]
- Lynch CJ, Adams SH: Branched-chain amino acids in metabolic signalling and insulin resistance. Nat Rev Endocrinol. 2014 Dec;10(12):723-36. doi: 10.1038/nrendo.2014.171. Epub 2014 Oct 7. [PubMed:25287287 ]
- Xiao F, Yu J, Guo Y, Deng J, Li K, Du Y, Chen S, Zhu J, Sheng H, Guo F: Effects of individual branched-chain amino acids deprivation on insulin sensitivity and glucose metabolism in mice. Metabolism. 2014 Jun;63(6):841-50. doi: 10.1016/j.metabol.2014.03.006. Epub 2014 Mar 15. [PubMed:24684822 ]
|
|---|